Knowledge Graph Construction with Claude
You have a pile of unstructured documents and need to answer questions that span them — "who works with people who worked on project X", "which vendors are connected to this incident". No single document contains the answer. RAG retrieval won't chain the facts for you. You need a knowledge graph: entities as nodes, typed relations as edges, so that multi-hop reasoning becomes graph traversal.
Building one used to mean training a named-entity recognizer on your domain, training a relation classifier, writing entity-resolution heuristics, and maintaining all three as your data shifted. With Claude, each of those stages becomes a prompt.
What you'll learn
By the end of this guide you will be able to:
- Use structured outputs to extract typed entities and subject–predicate–object triples from arbitrary text with no training data
- Apply Claude-driven entity resolution to collapse surface-form variants into canonical nodes, replacing brittle string-similarity heuristics
- Assemble and query an in-memory graph, and run multi-hop questions by serializing subgraphs back to Claude
- Measure extraction quality with precision/recall against a gold set and reason about the cost/quality tradeoff between Haiku and Sonnet
Everything runs in memory with no database. The techniques transfer directly to Neo4j, Neptune, or a Postgres adjacency table when you need to scale.
Prerequisites
- Python 3.11+
- Anthropic API key (get one here)
- Basic familiarity with graphs (nodes, edges, traversal)
Setup
%%capture
%pip install anthropic requests networkx matplotlib python-dotenv pydanticimport json
from collections import defaultdict
from pathlib import Path
from typing import Literal
from urllib.parse import quote
import anthropic
import matplotlib.pyplot as plt
import networkx as nx
import requests
from dotenv import load_dotenv
from pydantic import BaseModel
load_dotenv()
client = anthropic.Anthropic()
EXTRACTION_MODEL = "claude-haiku-4-5"
SYNTHESIS_MODEL = "claude-sonnet-4-6"We use two models. Haiku handles the high-volume, schema-constrained extraction work where speed and cost matter more than nuance. Sonnet handles entity resolution and summarization, where the model needs to weigh conflicting evidence across documents.
Building a corpus
We need a handful of documents that talk about overlapping entities, so that entity resolution has real work to do. The Apollo program is a good test bed: six short Wikipedia summaries that all mention NASA, the Moon, several astronauts, and a launch vehicle — but each article names them slightly differently.
We fetch summaries from the Wikipedia REST API rather than full articles to keep token costs low. For a production pipeline you would chunk full documents; the extraction logic is identical.
ARTICLE_TITLES = [
"Apollo program",
"Apollo 11",
"Neil Armstrong",
"Saturn V",
"Buzz Aldrin",
"Kennedy Space Center",
]
WIKI_API = "https://en.wikipedia.org/api/rest_v1/page/summary/"
HEADERS = {"User-Agent": "claude-cookbooks/1.0 (https://github.com/anthropics/claude-cookbooks)"}
def fetch_summary(title: str) -> str:
slug = quote(title.replace(" ", "_"), safe="")
r = requests.get(WIKI_API + slug, headers=HEADERS, timeout=10)
r.raise_for_status()
return r.json()["extract"]
documents = []
for i, title in enumerate(ARTICLE_TITLES):
try:
documents.append({"id": i, "title": title, "text": fetch_summary(title)})
except requests.RequestException as e:
print(f"Skipping {title}: {e}")
if not documents:
raise RuntimeError("No documents loaded — check network and Wikipedia API availability")
print(f"Loaded {len(documents)} documents\n")
print(f"Sample — {documents[0]['title']}:\n{documents[0]['text'][:300]}...")Loaded 6 documents Sample — Apollo program: The Apollo program, also known as Project Apollo, was the United States human spaceflight program led by NASA, which landed the first humans on the Moon in 1969. Apollo was conceived during Project Mercury and executed after Project Gemini. It was conceived in 1960 as a three-person spacecraft durin...
Entity and relation extraction
Classical NER tags spans of text with labels (PERSON, ORG, LOC). Classical relation extraction then classifies pairs of spans into relation types. Both traditionally require labeled training data per domain.
We collapse both stages into a single Claude call per document. The key is structured outputs: we define the output shape as a Pydantic model and pass it to client.messages.parse(). Claude's response is guaranteed to validate against that schema and comes back as a typed Python object — no regex parsing, no JSON decode errors, no defensive isinstance checks.
EntityType = Literal["PERSON", "ORGANIZATION", "LOCATION", "EVENT", "ARTIFACT"]
ENTITY_TYPES = ["PERSON", "ORGANIZATION", "LOCATION", "EVENT", "ARTIFACT"]
class Entity(BaseModel):
name: str
type: EntityType
description: str
class Relation(BaseModel):
source: str
predicate: str
target: str
class ExtractedGraph(BaseModel):
entities: list[Entity]
relations: list[Relation]
EXTRACTION_PROMPT = """Extract a knowledge graph from the document below.
<document>
{text}
</document>
Guidelines:
- Extract only entities that are central to what this document is about — skip incidental mentions.
- For each entity, write a one-sentence description grounded in this document. These descriptions are used later to disambiguate entities with similar names.
- Predicates should be short verb phrases ("commanded", "launched from", "part of").
- Every relation must connect two entities you extracted."""
def extract(text: str) -> ExtractedGraph:
response = client.messages.parse(
model=EXTRACTION_MODEL,
max_tokens=2048,
messages=[{"role": "user", "content": EXTRACTION_PROMPT.format(text=text)}],
output_format=ExtractedGraph,
)
return response.parsed_outputraw_entities = []
raw_relations = []
for doc in documents:
try:
result = extract(doc["text"])
except anthropic.APIError as e:
print(f"Skipping {doc['title']}: {e}")
continue
for ent in result.entities:
raw_entities.append({**ent.model_dump(), "source_doc": doc["title"]})
for rel in result.relations:
raw_relations.append({**rel.model_dump(), "source_doc": doc["title"]})
print(
f"{doc['title']:<25} {len(result.entities):>3} entities {len(result.relations):>3} relations"
)
print(f"\nTotal: {len(raw_entities)} raw entities, {len(raw_relations)} raw relations")Apollo program 8 entities 7 relations Apollo 11 6 entities 5 relations Neil Armstrong 3 entities 2 relations Saturn V 5 entities 4 relations Buzz Aldrin 6 entities 6 relations Kennedy Space Center 8 entities 10 relations Total: 36 raw entities, 34 raw relations
Let's look at what was extracted. Notice how the same real-world entity appears under different surface forms across documents — this is the entity resolution problem we solve next.
by_type = defaultdict(list)
for e in raw_entities:
by_type[e["type"]].append(e["name"])
for etype, names in sorted(by_type.items()):
print(f"{etype} ({len(names)}):")
for name in sorted(set(names)):
print(f" {name}")
print()ARTIFACT (3): Launch Complex 39 Saturn V Skylab EVENT (11): Apollo 11 Apollo program Gemini 12 Project Gemini Project Mercury Skylab program Space Shuttle program LOCATION (5): Merritt Island Moon ORGANIZATION (7): Cape Canaveral Space Force Station John F. Kennedy Space Center NASA U.S. Congress PERSON (10): Buzz Aldrin Dwight D. Eisenhower Edwin Aldrin Jim Lovell John F. Kennedy Michael Collins Neil Alden Armstrong Neil Armstrong
Entity resolution
The raw extraction gives us overlapping mentions: "NASA" and "National Aeronautics and Space Administration", "Neil Armstrong" and "Armstrong", possibly "the Moon" and "Moon". If we build a graph directly from this, we get a fractured mess where the same concept is split across disconnected nodes.
Traditional approaches use string similarity (edit distance, Jaccard on tokens) plus blocking rules. That works for typos but fails on "Edwin Aldrin" vs "Buzz Aldrin" — two names with zero character overlap that refer to the same person.
We instead ask Claude to cluster entities of each type, using the one-line descriptions from extraction as disambiguation context. The descriptions matter: "Armstrong — first person to walk on the Moon" and "Armstrong — jazz trumpeter" have the same name but should not merge.
class Cluster(BaseModel):
canonical: str
aliases: list[str]
class ResolvedClusters(BaseModel):
clusters: list[Cluster]
RESOLVE_PROMPT = """Below are {entity_type} entities extracted from several documents. Some are different surface forms of the same real-world entity.
<entities>
{entity_list}
</entities>
Cluster them. Each input name must appear in exactly one cluster's aliases list. Entities that are genuinely distinct get their own single-element cluster. Use the descriptions to avoid merging entities that merely share a name. The canonical name should be the most complete, unambiguous form."""
def resolve(entity_type: str, entities: list[dict]) -> list[Cluster]:
unique = {}
for e in entities:
unique.setdefault(e["name"], e["description"])
entity_list = "\n".join(f"- {name}: {desc}" for name, desc in unique.items())
response = client.messages.parse(
model=SYNTHESIS_MODEL,
max_tokens=2048,
messages=[
{
"role": "user",
"content": RESOLVE_PROMPT.format(entity_type=entity_type, entity_list=entity_list),
}
],
output_format=ResolvedClusters,
)
return response.parsed_output.clustersTwo failure modes to watch for. First, any raw name Claude leaves out of every cluster silently disappears from the graph, because alias_to_canonical has no entry for it — a production resolver should fall back to a single-element cluster for unmatched names so nothing is lost. Second, the resolver can over-merge: a specific mission like "Gemini 12" may get folded into the broader "Project Gemini" because the descriptions overlap. The first loses nodes, the second loses precision. Both are worth spot-checking in the output below.
alias_to_canonical = {}
canonical_info = {}
for etype in ENTITY_TYPES:
entities_of_type = [e for e in raw_entities if e["type"] == etype]
if not entities_of_type:
continue
try:
clusters = resolve(etype, entities_of_type)
except anthropic.APIError as e:
print(f"Resolve failed for {etype}: {e}; treating each name as its own cluster")
clusters = [
Cluster(canonical=n, aliases=[n]) for n in {x["name"] for x in entities_of_type}
]
for cluster in clusters:
canonical_info[cluster.canonical] = {"type": etype, "aliases": cluster.aliases}
for alias in cluster.aliases:
alias_to_canonical[alias] = cluster.canonical
before = len({e["name"] for e in raw_entities})
after = len(canonical_info)
print(f"Entity resolution: {before} unique names → {after} canonical entities\n")
for canonical, info in sorted(canonical_info.items()):
aliases = [a for a in info["aliases"] if a != canonical]
alias_str = f" (also: {', '.join(aliases)})" if aliases else ""
print(f"{info['type']:<14} {canonical}{alias_str}")Entity resolution: 24 unique names → 22 canonical entities EVENT Apollo 11 EVENT Apollo program PERSON Buzz Aldrin (also: Edwin Aldrin) ORGANIZATION Cape Canaveral Space Force Station PERSON Dwight D. Eisenhower EVENT Gemini 12 PERSON Jim Lovell PERSON John F. Kennedy ORGANIZATION John F. Kennedy Space Center ARTIFACT Launch Complex 39 LOCATION Merritt Island PERSON Michael Collins LOCATION Moon ORGANIZATION NASA PERSON Neil Alden Armstrong (also: Neil Armstrong) EVENT Project Gemini EVENT Project Mercury ARTIFACT Saturn V ARTIFACT Skylab EVENT Skylab program EVENT Space Shuttle program ORGANIZATION U.S. Congress
Assembling the graph
With a clean alias map, we rewrite every relation endpoint to its canonical form and load the result into NetworkX. We use a MultiDiGraph because two entities can be connected by several distinct predicates ("launched from" and "operated by"), and direction matters ("Armstrong commanded Apollo 11" is not the same edge as "Apollo 11 commanded Armstrong").
Each node carries its type, the set of documents that mention it, and a mention count. Each edge carries its predicate and source document.
G = nx.MultiDiGraph()
for e in raw_entities:
canonical = alias_to_canonical.get(e["name"])
if canonical is None:
continue
if canonical not in G:
G.add_node(
canonical,
type=canonical_info[canonical]["type"],
description=e["description"],
source_docs=[],
mentions=0,
)
G.nodes[canonical]["source_docs"].append(e["source_doc"])
G.nodes[canonical]["mentions"] += 1
for r in raw_relations:
src = alias_to_canonical.get(r["source"])
tgt = alias_to_canonical.get(r["target"])
if src and tgt and src != tgt:
G.add_edge(src, tgt, predicate=r["predicate"], source_doc=r["source_doc"])
for n in G.nodes:
G.nodes[n]["source_docs"] = sorted(set(G.nodes[n]["source_docs"]))
print(f"Graph: {G.number_of_nodes()} nodes, {G.number_of_edges()} edges")
print(f"Connected components: {nx.number_weakly_connected_components(G)}")
print("\nMost connected entities:")
for node, deg in sorted(G.degree(), key=lambda x: -x[1])[:5]:
print(f" {node:<35} degree {deg:>2} ({G.nodes[node]['type']})")Graph: 22 nodes, 34 edges Connected components: 1 Most connected entities: Apollo program degree 9 (EVENT) Apollo 11 degree 9 (EVENT) John F. Kennedy Space Center degree 7 (ORGANIZATION) Neil Alden Armstrong degree 5 (PERSON) Buzz Aldrin degree 5 (PERSON)
COLOR = {
"PERSON": "#4e79a7",
"ORGANIZATION": "#f28e2c",
"LOCATION": "#76b7b2",
"EVENT": "#e15759",
"ARTIFACT": "#af7aa1",
}
plt.figure(figsize=(14, 10))
pos = nx.spring_layout(G, k=1.5, seed=42)
node_colors = [COLOR[G.nodes[n]["type"]] for n in G.nodes]
node_sizes = [300 + 200 * G.degree(n) for n in G.nodes]
nx.draw_networkx_nodes(G, pos, node_color=node_colors, node_size=node_sizes, alpha=0.9)
nx.draw_networkx_labels(G, pos, font_size=8)
nx.draw_networkx_edges(G, pos, alpha=0.3, arrows=True, arrowsize=10)
handles = [
plt.Line2D([0], [0], marker="o", color="w", markerfacecolor=c, markersize=10, label=t)
for t, c in COLOR.items()
if any(G.nodes[n]["type"] == t for n in G.nodes)
]
plt.legend(handles=handles, loc="upper left")
plt.title("Apollo Program Knowledge Graph")
plt.axis("off")
plt.tight_layout()
plt.show()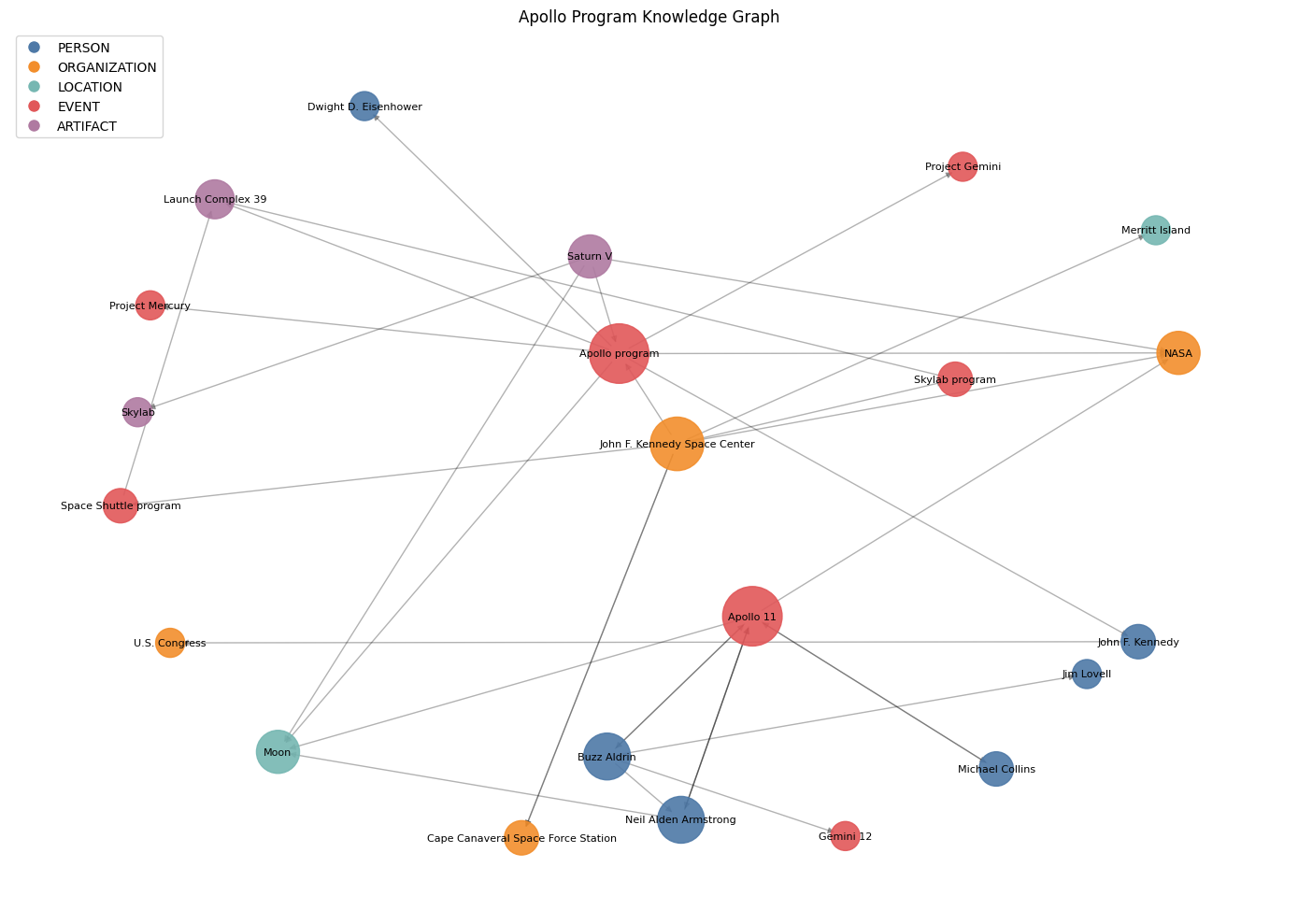
Node size scales with degree — the hubs are the entities that tie the corpus together. Color encodes type: if your graph is mostly one color, your corpus is narrow; a good mix means the extractor is finding the full cast of people, places, and things. A single connected component means entity resolution did its job — fragmented islands would indicate variants that should have merged but didn't.
Entity summarization
Each node currently carries only the one-line description from whichever document mentioned it first. For the hub nodes — the ones that show up in many documents — we can do much better: pool every mention, add the graph neighborhood as context, and have Claude synthesize a proper profile.
This is the step that turns a graph of labels into a graph of knowledge. The summaries become the node content you surface in search results or feed to downstream QA.
class TimeRange(BaseModel):
start: str # YYYY or YYYY-MM, or "unknown"
end: str # YYYY or YYYY-MM, or "ongoing"
class EntityProfile(BaseModel):
summary: str
key_facts: list[str]
time_range: TimeRange
SUMMARIZE_PROMPT = """Generate a knowledge-graph profile for this entity.
Entity: {name} ({etype})
Source excerpts mentioning this entity:
{excerpts}
Known relations in the graph:
{relations}
Write a 2-3 paragraph factual summary synthesized from the excerpts, resolving any contradictions by preferring the most specific claim. Include 3-5 atomic key facts, each traceable to the sources. For the time range, use YYYY or YYYY-MM format, or "unknown"/"ongoing" where appropriate. Do not invent facts not supported by the excerpts."""
def summarize_entity(name: str) -> EntityProfile:
# Reads module-level G and documents built earlier in the notebook.
docs_with_entity = G.nodes[name]["source_docs"]
excerpts = "\n\n".join(
f"[{d['title']}]\n{d['text']}" for d in documents if d["title"] in docs_with_entity
)
relations = (
"\n".join(
f"- {name} --{d['predicate']}--> {tgt}" for _, tgt, d in G.out_edges(name, data=True)
)
+ "\n"
+ "\n".join(
f"- {src} --{d['predicate']}--> {name}" for src, _, d in G.in_edges(name, data=True)
)
)
response = client.messages.parse(
model=SYNTHESIS_MODEL,
max_tokens=1500,
messages=[
{
"role": "user",
"content": SUMMARIZE_PROMPT.format(
name=name, etype=G.nodes[name]["type"], excerpts=excerpts, relations=relations
),
}
],
output_format=EntityProfile,
)
return response.parsed_outputhub_nodes = [n for n, _ in sorted(G.degree(), key=lambda x: -x[1])[:3]]
for node in hub_nodes:
profile = summarize_entity(node)
G.nodes[node]["profile"] = profile.model_dump()
print(f"═══ {node} ═══")
print(profile.summary)
print(f"\nTime range: {profile.time_range.start} – {profile.time_range.end}")
print("Key facts:")
for fact in profile.key_facts:
print(f" • {fact}")
print()═══ Apollo program ═══ The Apollo program, also known as Project Apollo, was a United States human spaceflight program led by NASA, primarily aimed at landing humans on the Moon. It was conceived in 1960 during the presidency of Dwight D. Eisenhower as a three-person spacecraft program, building upon the foundations laid by Project Mercury. The program was later formally dedicated to President John F. Kennedy's national goal, announced before the U.S. Congress on May 25, 1961, of landing a man on the Moon and returning him safely to Earth. Apollo followed Project Gemini in NASA's sequential human spaceflight efforts. The Apollo program successfully landed the first humans on the Moon in 1969. It relied on the Saturn V, a super heavy-lift launch vehicle developed by NASA specifically for the program, which flew from 1967 to 1973 and supported nine crewed flights to the Moon. Launch operations were conducted from Launch Complex 39 at Kennedy Space Center on Merritt Island, Florida, which has served as NASA's primary launch center since 1968. Time range: 1960 – 1973 Key facts: • The Apollo program was conceived in 1960 under the presidency of Dwight D. Eisenhower as a three-person spacecraft program. • Apollo was dedicated to President John F. Kennedy's goal of landing a man on the Moon, as stated in his address to Congress on May 25, 1961. • The Apollo program successfully landed the first humans on the Moon in 1969. • The Saturn V rocket, developed under the Apollo program, flew from 1967 to 1973 and supported nine crewed lunar flights. • Launch operations for the Apollo program were carried out from Kennedy Space Center Launch Complex 39. ═══ Apollo 11 ═══ Apollo 11 was the American spaceflight that achieved the historic first crewed lunar landing, representing the fifth crewed mission of NASA's Apollo program. The mission was commanded by Neil Armstrong, with Michael Collins serving as Command Module Pilot and Edwin "Buzz" Aldrin as Lunar Module Pilot. All three crew members were on their second and final spaceflights. During the mission, Neil Armstrong became the first person to walk on the Moon, followed by Buzz Aldrin, making him the second. Michael Collins remained in the Command Module in lunar orbit. The mission stands as a landmark achievement in human spaceflight history. As of 2025, Buzz Aldrin is the last surviving crew member of Apollo 11, following the deaths of Armstrong in 2012 and Collins in 2021. Time range: 1969 – 1969 Key facts: • Apollo 11 was the fifth crewed mission of NASA's Apollo program and the first to land humans on the Moon. • The crew consisted of Commander Neil Armstrong, Command Module Pilot Michael Collins, and Lunar Module Pilot Buzz Aldrin. • Neil Armstrong was the first person to walk on the Moon, with Buzz Aldrin becoming the second. • All three Apollo 11 crew members were on their second and final spaceflights. • Buzz Aldrin is the last surviving member of the Apollo 11 crew, following the deaths of Armstrong in 2012 and Collins in 2021. ═══ John F. Kennedy Space Center ═══ The John F. Kennedy Space Center (KSC) is a NASA field center situated on Merritt Island, Florida, on the east coast of the state. It is one of ten field centers operated by NASA and has served as the agency's primary launch center for American spaceflight, research, and technology since 1968. Launch operations for several major human spaceflight programs, including Apollo, Skylab, and the Space Shuttle, were conducted from KSC's Launch Complex 39 and managed by the center. KSC operates in close proximity to Cape Canaveral Space Force Station (CCSFS), with the two entities maintaining a collaborative relationship. Their management teams work closely together, share resources, and operate facilities on each other's property, reflecting a high degree of integration between the two installations. Time range: 1968 – ongoing Key facts: • John F. Kennedy Space Center is located on Merritt Island, Florida, and is one of ten NASA field centers. • KSC has been NASA's primary launch center for American spaceflight, research, and technology since 1968. • Launch operations for the Apollo, Skylab, and Space Shuttle programs were carried out from KSC's Launch Complex 39. • KSC is adjacent to Cape Canaveral Space Force Station, and the two entities share resources and operate facilities on each other's property.
Querying the graph
The payoff of building a knowledge graph is multi-hop reasoning: answering questions that require chaining facts that never co-occur in a single document. "Which locations are connected to people who flew on Apollo 11?" needs the extractor to have found person→mission edges in one document and person→location edges in another, then the resolver to have unified the person nodes so those edges actually meet.
We serialize a relevant subgraph as triples and let Claude reason over it. For comparison, we first ask the same question with no graph context.
def serialize_subgraph(center: str, hops: int = 2) -> str:
nodes = {center}
frontier = {center}
for _ in range(hops):
nxt = set()
for n in frontier:
nxt |= set(G.successors(n)) | set(G.predecessors(n))
frontier = nxt - nodes
nodes |= frontier
sub = G.subgraph(nodes)
lines = [f"({s}) --[{d['predicate']}]--> ({t})" for s, t, d in sub.edges(data=True)]
return "\n".join(sorted(set(lines)))
def ask(question: str, graph_context: str | None = None) -> str:
if graph_context is not None:
prompt = f"""Answer using only the knowledge graph below. Cite the specific edges that support your answer.
<graph>
{graph_context}
</graph>
Question: {question}"""
else:
prompt = question
response = client.messages.create(
model=SYNTHESIS_MODEL,
max_tokens=500,
messages=[{"role": "user", "content": prompt}],
)
text_block = next((b for b in response.content if b.type == "text"), None)
if text_block is None:
raise ValueError(f"No text block in response (stop_reason={response.stop_reason})")
return text_block.textcenter = next((n for n in G.nodes if "Apollo" in n), hub_nodes[0])
print(f"Querying 2-hop neighborhood of: {center}\n")
subgraph = serialize_subgraph(center, hops=2)
question = "Which locations are connected to people who were part of Apollo 11, and how?"
print("WITHOUT graph context:")
print(ask(question))
print("\n" + "─" * 60 + "\n")
print("WITH graph context:")
print(ask(question, subgraph))Querying 2-hop neighborhood of: Apollo program WITHOUT graph context: # Locations Connected to Apollo 11 Personnel ## Neil Armstrong (Commander) - **Wapakoneta, Ohio** – birthplace and hometown; the **Neil Armstrong Air & Space Museum** is located there - **Purdue University, Indiana** – where he studied aeronautical engineering - **Edwards Air Force Base, California** – where he worked as a test pilot - **Cincinnati, Ohio** – where he lived later in life and died (2012) ## Buzz Aldrin (Lunar Module Pilot) - **Montclair, New Jersey** – birthplace - **West Point, New York** – attended the U.S. Military Academy - **MIT, Massachusetts** – earned his doctorate in astronautics - **Los Angeles area** – longtime residence after NASA career ## Michael Collins (Command Module Pilot) - **Rome, Italy** – birthplace (son of a U.S. Army officer) - **Washington, D.C.** – attended school; later directed the **Smithsonian National Air and Space Museum** - **West Point** – attended the Military Academy ## Shared/Mission Locations - **Kennedy Space Center, Florida** – launch site (July 16, 1969) - **Houston, Texas** – Mission Control and astronaut training (Johnson Space Center) - **Pacific Ocean** – splashdown and recovery site - **Honolulu, Hawaii** – where the crew was initially quarantined aboard the USS Hornet Would you like more detail on any individual? ──────────────────────────────────────────────────────────── WITH graph context: ## Locations Connected to Apollo 11 Personnel Based on the knowledge graph, here is what can be determined about people connected to Apollo 11 and their location relationships: ### Neil Alden Armstrong Neil Armstrong is the person directly identified as part of Apollo 11, supported by: - **(Apollo 11) --[commanded by]--> (Neil Alden Armstrong)** - **(Neil Alden Armstrong) --[commanded]--> (Apollo 11)** He is connected to the following location: #### 🌕 The Moon - **(Neil Alden Armstrong) --[walked on]--> (Moon)** This is the only **direct** person-to-location edge in the graph for an Apollo 11 crew member. --- ### Indirect Location Connection via Apollo 11 itself: Apollo 11 as a mission is also connected to the Moon: - **(Apollo 11) --[landed on]--> (Moon)** --- ### Limitations The knowledge graph does **not** include location data for any other Apollo 11 crew members (e.g., Buzz Aldrin or Michael Collins), so no connections can be drawn for them. The only person-location relationship supported by the graph is **Neil Armstrong → walked on → the Moon**.
The ungrounded answer draws on Claude's pretraining and may be correct — Apollo 11 is famous. But the grounded answer is traceable: every claim cites an edge we extracted from a specific document. On a private corpus where Claude has no prior knowledge, only the grounded answer works at all.
Evaluation
Knowledge graph quality is measured with precision and recall against a gold set. We ship a small hand-labeled set in data/sample_triples.json covering two of the articles, plus data/alias_map.json which normalizes surface-form variants to the gold names so that "the Moon" and "Moon" count as the same hit.
The check below scores two things side by side: raw extractor output, and the same entities after passing through alias_to_canonical from the resolution step. When the resolver picks a canonical form the alias map knows about (or that matches gold verbatim), resolved recall climbs. When it picks a verbose form the alias map doesn't cover — say "Neil Alden Armstrong" — resolved recall can drop, because a name that matched gold before resolution no longer does after. That's not a resolver bug; it's a scoring artifact. The fix is to extend alias_map.json whenever you see a canonical form the scorer doesn't recognize.
This cell scores entities only. The standalone script also scores relations, matching on (source, target) pairs with predicate wording ignored — so its relation recall is an upper bound. Run it from the repo root:
uv run python capabilities/knowledge_graph/evaluation/eval_extraction.py# Expects the kernel launched from this notebook's directory
# (capabilities/knowledge_graph/). Falls back to repo root.
data_dir = Path("data")
if not data_dir.exists():
data_dir = Path("capabilities/knowledge_graph/data")
with open(data_dir / "sample_triples.json", encoding="utf-8") as f:
gold = json.load(f)
with open(data_dir / "alias_map.json", encoding="utf-8") as f:
ALIASES = json.load(f)
def norm(name: str) -> str:
lower = name.lower().strip()
return ALIASES.get(lower, lower)
def prf(predicted: set, gold: set) -> tuple[float, float, float]:
tp = len(predicted & gold)
p = tp / len(predicted) if predicted else 0.0
r = tp / len(gold) if gold else 0.0
f1 = 2 * p * r / (p + r) if (p + r) else 0.0
return p, r, f1
print("Raw extraction vs resolved-graph recall against gold:\n")
for doc_title, labels in gold.items():
gold_names = {norm(e["name"]) for e in labels["entities"]}
raw = {norm(e["name"]) for e in raw_entities if e["source_doc"] == doc_title}
rp, rr, rf = prf(raw, gold_names)
resolved = {
norm(alias_to_canonical.get(e["name"], e["name"]))
for e in raw_entities
if e["source_doc"] == doc_title
}
_, resolved_r, _ = prf(resolved, gold_names)
print(f"{doc_title:<20} raw F1={rf:.2f} (P={rp:.2f} R={rr:.2f}) resolved R={resolved_r:.2f}")
missed = gold_names - resolved
if missed:
print(f" still missed after resolution: {', '.join(sorted(missed))}")Raw extraction vs resolved-graph recall against gold: Apollo 11 raw F1=0.71 (P=1.00 R=0.55) resolved R=0.55 still missed after resolution: apollo program, columbia, kennedy space center, lunar module eagle, saturn v Neil Armstrong raw F1=0.55 (P=1.00 R=0.38) resolved R=0.38 still missed after resolution: gemini 8, korean war, nasa, purdue university, united states navy
Scaling up
This notebook processed six documents in memory. Production knowledge graphs are built from thousands. A few notes on scaling:
Extraction cost. Haiku is cheap enough to run on large corpora, but prompt caching cuts costs further when your extraction schema and instructions stay fixed — cache the system prompt and schema, pay full price only for the document text. The Message Batches API gives 50% off for jobs that can wait up to 24 hours.
Entity resolution at scale. Feeding ten thousand PERSON entities to Claude in one prompt doesn't work. Block first: group candidates by cheap signals (same last name, overlapping tokens, embedding similarity) so Claude only arbitrates within small blocks. The resolution prompt above works unchanged on blocks of 50–100.
Incremental updates. When a new document arrives, extract its entities, resolve them against the existing canonical set (not against each other), and add only the new edges. Re-summarize an entity only when its source-document set changes materially.
Storage. NetworkX is fine to a few hundred thousand edges. Beyond that, the schema maps directly onto a property graph (Neo4j, Neptune) or three Postgres tables: entities(id, name, type, summary), relations(source_id, target_id, predicate), aliases(entity_id, alias). The extraction and resolution code doesn't change — only the persistence layer does.
Summary
You've built a complete knowledge graph pipeline with nothing but prompts:
- Extraction — a single structured-output call per document replaced what used to be a trained NER model plus a trained relation classifier. The Pydantic schema is the only "training" required.
- Resolution — Claude clustered surface forms using the extraction descriptions as context, catching cases like "Edwin Aldrin" → "Buzz Aldrin" that string similarity misses entirely.
- Summarization — hub nodes got rich profiles synthesized across every document that mentioned them, with structured time ranges and traceable key facts.
- Querying — serialized subgraphs let Claude answer multi-hop questions with edge-level citations, grounding answers in the graph you built rather than pretraining.
The evaluation harness in evaluation/ gives you a feedback loop: change the extraction prompt, rerun the scorer, watch the F1 move. That loop is what turns a demo into a production system.
- Extracting structured JSON — the tool-use approach to the same extraction pattern, useful when you're already in an agentic tool-calling flow
- Retrieval augmented generation — the complementary approach when you need document retrieval rather than fact traversal
- Contextual embeddings — enriching chunks before embedding, the same "add context before indexing" idea applied to vector search
